This is probably not my last re-analysis of the same dataset, and I'd want to encourage others to use other phylogenetic methods on our alignments, or use our alignments to analyse their own SCNA sequences.
Our published study used both sequence-based phylogenetic data and genomic data to clear up the muddled relationships between different subtypes of voltage-gated sodium channels in vertebrates. Us humans, and most other mammals, have ten different types of these α subunits, encoded by genes of the SCNA family. The different subtypes have acquired different properties and are expressed in different parts of the nervous system and some other tissues in the body where electrical signaling is necessary. One of the subtypes, the product of the SCN7A gene, has even evolved to such a degree that it's not a channel anymore. It's probably more like a sodium "sensor". There are also specific diseases, different types of epilepsy, seizures and paralyses, that are associated with mutations of the various subtypes.
The same gene family in true bony fishes (teleosts) encodes up to eight different subtypes, almost as many as the mammalian family. The funny evolutionary twist is that we have evolved almost the same amount of subtypes by completely different mechanisms! In tetrapods, the fleshy-limbed mostly terrestrial group that we belong to, local duplications of genes on the same chromosomes gave rise to new subtypes, while in teleost fishes it was their ancestral duplication of the whole genome that did the same on different chromosomes! You can see the difference in the image below.
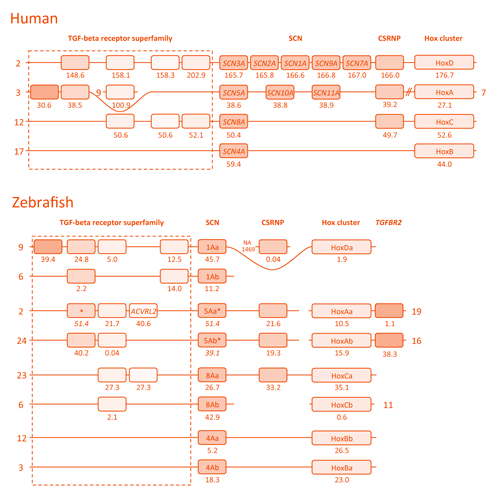
Chromosome organization of the SCNA genes and some of their neighboring genes in humans and zebrafish. Each numbered line represents a different chromosome. The ten human genes are named in a series from SCN1A to SCN11A, skipping the number 6. The duplicates in teleost fish are denoted by the letters "a" and "b".
This means that when doing comparative studies, it's important to know that while all vertebrate voltage-gated sodium channels have a common origin, they're not always directly related to each other.
Using more powerful and careful methods than we'd used before, I made new evolutionary trees that agree even better with these conclusions. Most of all these new trees are better and clearer examples of the evolutionary relationships between different voltage-gated sodium channel subtypes. You can see figures of the new trees and access all the source files and datasets in Figshare.
Phylogenetic maximum likelihood analyses of the voltage-gated sodium channel α subunit (SCNα) gene family based on amino acid sequence alignments. The sequences and alignments described in Widmark et. al. (2011) Molecular Biology and Evolution 28(1):859-71 (1) were used to re-analyze the phylogenetic relationships of vertebrate SCNα subtypes with more powerful methods.
While our previous study solved the evolutionary relationships between different SCNA subtypes, we had to rely on the genomic data on the chromosome locations of the genes to solve some inconsistencies in the original phylogenetic trees. Our study was the first to do phylogenetic analyses from alignments of the whole α subunit sequences, which are over 2000 amino acids long! This work was almost entirely carried out by my colleague Jenny Widmark and our supervisor Dan Larhammar. The fact that most previous studies had only analysed parts of the proteins was the reason they had reached the wrong conclusions about how different subunits were related.
Assembling and aligning all that sequence data from the genomic sequences of multiple vertebrate species took a lot of time and thought, so I thought it would be a shame not to use it to find out if better methods could solve what the published tree couldn't and if they could provide more and better evidence for our conclusions, especially when all the different analyses are considered together.
As a summary, I put together this tree over the relationships between the different human subtypes. As you can see, the positions of the different subtypes in the trees agree with the chromosome locations.
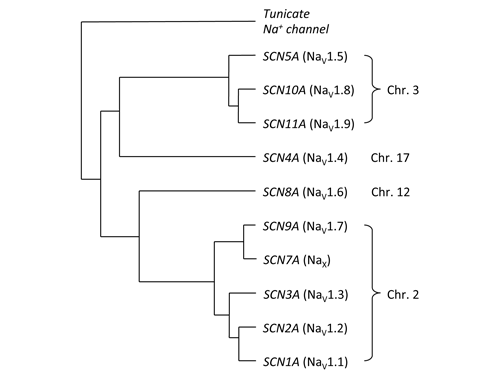
The most up-to-date view of the relationship between the different human voltage-gated sodium channel subtypes (the names of the proteins are in parenthesis), based on my latest re-analyses. Teleost fish have duplicates of SCN4A, SCN8A and the genes that gave rise to SCN5A, -10A and -11A and SCN1A, -2A, -3A, -9A and -7A respectively.
Previous analyses put some of the more divergent subtypes, like the products of the SCN11A and SCN7A genes at the base of the tree when they are in fact some of the newest subtypes! Now we can also say that SCN4A and the SCN5A, -10A, -11A-group are more closely related, and that SCN8A and the SCN1A, -2A, -3A, -9A, -7A-group are more closely related. You can see some the older analyses that don't actually give the correct evolutionary view displayed on IUPHAR's* database of receptors and ion channels (link). Another example is displayed in Wikipedia (link).
The next step is to contact IUPHAR and ask them to review our new data so that our updated view of the voltage-gated sodium channels reaches all the people it potentially benefits, everyone doing functional research on sodium channels. To do that I think an open science approach is clearly the best way to go.
* The International Union of Basic and Clinical Pharmacology decides on "official" receptor and ion channel nomenclature, among other things.
Widmark, J., Sundstrom, G., Ocampo Daza, D., & Larhammar, D. (2010). Differential Evolution of Voltage-Gated Sodium Channels in Tetrapods and Teleost Fishes Molecular Biology and Evolution, 28 (1), 859-871 DOI: 10.1093/molbev/msq257
No comments:
Post a Comment
Note: Only a member of this blog may post a comment.